Hello!
I am working on the SVI implementation of the hierarchical Poisson factorization (http://www.cs.columbia.edu/~blei/fogm/2020F/readings/GopalanHofmanBlei2015.pdf).
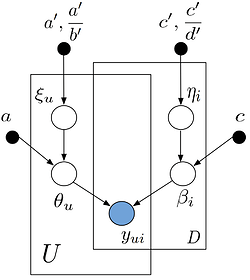
The structure of the model is as follows:
I have a pyro implementation of this model, but now I need it to be optimized for large sparse matrices, and I am primarily focusing on memory optimization. However, as far as I understand, pyro does not support sampling sparse tensors other than iterating all the elements one-by-one in a ‘for’ loop.
What is currently the best way to do SVI on sparse matrices? Is there a way to do that more or less efficient in pyro/numpyro?
I would really appreciate some advice!
what precisely is sparse? y_ui?
yes, exactly, my matrix of observations. I only want to sample the non-zero elements of it
probably your best bet would be to construct a custom my_log_prob and use it in a factor statement:
pyro.factor("obs", my_log_prob)
you can then do whatever custom sparse computation you want in my_log_prob. if the factor statement is inside of plate contexts it would need zero entries where there is no data. otherwise you could place the factor statement outside of plates
pyro.factor("obs", my_log_prob.sum())
taking care to scale the factor appropriately by the right multiplicative factor if you’re doing data subsampling (this scaling is otherwise handled automatically by the plates, but not if you “hide” the factor statement outside of the plate contexts)
1 Like