Hello numpyro users,
This post is about a class I created to build large models on numpyro through a class structure rather than the traditional pyro/numypro model function. I found this class to be very helpful in my personal project so I want to share it here and see if anyone else would be interested in using it. If there is interest let me know and I’ll submit a feature request on the numpyro github or share a personal repositiory. Also, this is very much a work in progress and I would love any and all feedback.
Background:
For some background I started working on a project a few months ago with pyro and then switched to numpyro because the model I was working with grew to a size that made numpyro’s speed appealing. My team and I found that as these models grow they get complicated quickly. We decided that it would be worth the time to create a class structure that could programmatically create numpyro models in order to cut down on lines of code while also keeping better track of the small details in the model.
Explanation of Example:
I will show a side by side comparison of this class structure and base numpyro’s way of creating a model. I’m just going to use my current project as an example because I think it shows off the model generation class use case well.
Here is a little background into our project, but the main thing I want to show off is how I create models so I will be brief.
We’re working with anthropologists to research ways in which we can accurately model the time since death, or postmortem interval (PMI). It will hopefully aid in the identification of an unknown individual and help to reconstruct the events around the time of death. We use numpyro to infer coefficients I labeled as β in the graph below. We go about this by observing the other variables and using MCMC to approximate β. COVARS represents characteristics of a body like body size, age, gender, etc. DECOMPS represents the decomposition characteristics on the body like skeletonization, mummification, etc.
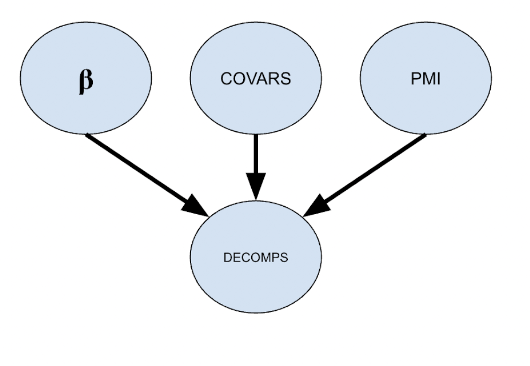
We then use the newly learned coefficients, observe covariates, and observe decomposition characteristics to make inferences about PMI. As a note, PMI has an exponential distribution so we are making inferences on log(PMI+1) so that we can use a normal distribution instead.
Code:
Using base numpyro to generate the model:
Feel free to glance through this! The main idea is this model long and hard to search through. It can be tedious in a research setting where a lot of changes are being made.
def prior_model(num_samples):
with seed(rng_seed=random.PRNGKey(0)):
#Age at death
livor_mortis_fixed_age_at_death_coefficient = numpyro.sample('livor_mortis_fixed_age_at_death_coefficient',dist.Normal(np.array([0, 0, 0, 0, 0]), 2).to_event(1))
skin_slippage_age_at_death_coefficient = numpyro.sample('skin_slippage_age_at_death_coefficient',dist.Normal(np.array([0, 0, 0, 0, 0]), 2).to_event(1))
marbling_age_at_death_coefficient = numpyro.sample('marbling_age_at_death_coefficient',dist.Normal(np.array([0, 0, 0, 0, 0]), 2).to_event(1))
#body size
livor_mortis_fixed_body_size_coefficient = numpyro.sample('livor_mortis_fixed_body_size_coefficient',dist.Normal(np.array([0, 0, 0]), 2).to_event(1))
skin_slippage_body_size_coefficient = numpyro.sample('skin_slippage_body_size_coefficient',dist.Normal(np.array([0, 0, 0]), 2).to_event(1))
marbling_body_size_coefficient = numpyro.sample('marbling_body_size_coefficient',dist.Normal(np.array([0, 0, 0]), 2).to_event(1))
#clothing
livor_mortis_fixed_clothing_coefficient = numpyro.sample('livor_mortis_fixed_clothing_coefficient',dist.Normal(np.array([0, 0, 0]), 2).to_event(1))
skin_slippage_clothing_coefficient = numpyro.sample('skin_slippage_clothing_coefficient',dist.Normal(np.array([0, 0, 0]), 2).to_event(1))
marbling_clothing_coefficient = numpyro.sample('marbling_clothing_coefficient',dist.Normal(np.array([0, 0, 0]), 2).to_event(1))
#is hanging
livor_mortis_fixed_is_hanging_coefficient = numpyro.sample('livor_mortis_fixed_is_hanging_coefficient',dist.Normal(0, 2))
skin_slippage_is_hanging_coefficient = numpyro.sample('skin_slippage_is_hanging_coefficient',dist.Normal(0, 2))
marbling_is_hanging_coefficient = numpyro.sample('marbling_is_hanging_coefficient',dist.Normal(0, 2))
#trauma that breaks the skin
livor_mortis_fixed_trauma_br_skin_coefficient = numpyro.sample('livor_mortis_fixed_trauma_br_skin_coefficient',dist.Normal(np.array([0, 0, 0, 0]), 2).to_event(1))
skin_slippage_trauma_br_skin_coefficient = numpyro.sample('skin_slippage_trauma_br_skin_coefficient',dist.Normal(np.array([0, 0, 0, 0]), 2).to_event(1))
marbling_trauma_br_skin_coefficient = numpyro.sample('marbling_trauma_br_skin_coefficient',dist.Normal(np.array([0, 0, 0, 0]), 2).to_event(1))
#intercept 1
livor_mortis_fixed_intercept = numpyro.sample('livor_mortis_fixed_intercept',dist.Normal(0,2))
skin_slippage_intercept = numpyro.sample('skin_slippage_intercept',dist.Normal(0,2))
marbling_intercept = numpyro.sample('marbling_intercept',dist.Normal(0,2))
#intercept 2
livor_mortis_fixed_logpi_intercept = numpyro.sample('livor_mortis_fixed_logpi_intercept',dist.Normal(0,2))
skin_slippage_logpi_intercept = numpyro.sample('skin_slippage_logpi_intercept',dist.Normal(0,2))
marbling_logpi_intercept = numpyro.sample('marbling_logpi_intercept',dist.Normal(0,2))
with plate('data', num_samples):
#covariates
age_at_death = numpyro.sample('age_at_death',dist.Categorical(logits=np.array([-4.239208520926547, -5.28994908412601, 3.846738437837947, -7.0859014343304585, -7.0859014343304585])))
body_size = numpyro.sample('body_size', dist.Categorical(logits=np.array([-2.2897760975840273, 0.9274377647576987, -1.4404764504491006])))
clothing = numpyro.sample('clothing',dist.Categorical(logits=np.array([0.39641516047,-1.623551043104,-1.166665718666])))
is_hanging = numpyro.sample('is_hanging',dist.Bernoulli(probs=0.00585284))
trauma_br_skin = numpyro.sample('trauma_br_skin',dist.Categorical(logits=np.array([-4.510859481371141, 0.09036250054334112, -0.2047254443224715, -4.0245010622075785])))
PMI = numpyro.sample('PMI',dist.Normal(2.1126173, 1.7476393))
#marbing
marbling_logpi = marbling_age_at_death_coefficient.at[...,age_at_death].get()
marbling_logpi += marbling_body_size_coefficient.at[...,body_size].get()
marbling_logpi += marbling_clothing_coefficient.at[...,clothing].get()
marbling_logpi += marbling_is_hanging_coefficient*is_hanging
marbling_logpi += marbling_trauma_br_skin_coefficient.at[...,trauma_br_skin].get()
marbling_logpi += marbling_intercept
marbling_logpi = PMI*marbling_logpi+marbling_logpi_intercept
marbling = numpyro.sample('marbling',dist.Bernoulli(logits=marbling_logpi))
#livor mortis fixed
livor_mortis_fixed_logpi = livor_mortis_fixed_age_at_death_coefficient.at[...,age_at_death].get()
livor_mortis_fixed_logpi += livor_mortis_fixed_body_size_coefficient.at[...,body_size].get()
livor_mortis_fixed_logpi += livor_mortis_fixed_clothing_coefficient.at[...,clothing].get()
livor_mortis_fixed_logpi += livor_mortis_fixed_is_hanging_coefficient*is_hanging
livor_mortis_fixed_logpi += livor_mortis_fixed_trauma_br_skin_coefficient.at[...,trauma_br_skin].get()
livor_mortis_fixed_logpi += livor_mortis_fixed_intercept
livor_mortis_fixed_logpi = PMI*livor_mortis_fixed_logpi+livor_mortis_fixed_logpi_intercept
livor_mortis_fixed = numpyro.sample('livor_mortis_fixed',dist.Bernoulli(logits=livor_mortis_fixed_logpi))
#skin_slippage
skin_slippage_logpi = skin_slippage_age_at_death_coefficient.at[...,age_at_death].get()
skin_slippage_logpi += skin_slippage_body_size_coefficient.at[...,body_size].get()
skin_slippage_logpi += skin_slippage_clothing_coefficient.at[...,clothing].get()
skin_slippage_logpi += skin_slippage_is_hanging_coefficient*is_hanging
skin_slippage_logpi += skin_slippage_trauma_br_skin_coefficient.at[...,trauma_br_skin].get()
skin_slippage_logpi += skin_slippage_intercept
skin_slippage_logpi = PMI*skin_slippage_logpi+skin_slippage_logpi_intercept
skin_slippage = numpyro.sample('skin_slippage',dist.Bernoulli(logits=skin_slippage_logpi))
return livor_mortis_fixed_age_at_death_coefficient,skin_slippage_age_at_death_coefficient,marbling_age_at_death_coefficient,livor_mortis_fixed_body_size_coefficient,\
skin_slippage_body_size_coefficient,marbling_body_size_coefficient,livor_mortis_fixed_clothing_coefficient,skin_slippage_clothing_coefficient,\
marbling_clothing_coefficient,livor_mortis_fixed_is_hanging_coefficient,skin_slippage_is_hanging_coefficient,marbling_is_hanging_coefficient,\
livor_mortis_fixed_trauma_br_skin_coefficient,skin_slippage_trauma_br_skin_coefficient,marbling_trauma_br_skin_coefficient, livor_mortis_fixed_intercept,\
skin_slippage_intercept,marbling_intercept,livor_mortis_fixed_logpi_intercept,skin_slippage_logpi_intercept,marbling_logpi_intercept,age_at_death,\
body_size,clothing,is_hanging,trauma_br_skin,PMI,marbling,livor_mortis_fixed,skin_slippage
Using class structure to generate the model:
This model generation class has three main components, .add_node(), .create_model(), and the “node_function”.
1. .add_node() and the node function:
The class has an emphasis on the graph representation of the model and is more intuitive if viewed from that perspective. We initially wanted to use a package like NetworkX to define the graph structure and still can, however I was only using .add_node() or .add_edge() in NetworkX so creating our own class structure seemed more straightforward.
The add_node function is how you add variables to a model. It takes in the name of that node and either a prior distribution or a “node_function”. The node function is just a function that returns the distribution of that node, and the parameters of that function will be used by the class to infer the parents of that node.
The idea of the node function is to provide the way in which a variable or “node” is calculated, but only if it depends on other variables. The model generation class will provide the arguments and the user returns the distribution. In my experience that allows for flexibility in the model similar to base pyro/numpyro, but it also cleans up code, cuts down on the number of lines, and allows for easy changes.
There are a few rules to the node function.
-
The parameters for a node function must exist as a node in the graph through .add_node() or through the arg_priors argument when passing a node function in .add_node(). Add_node and arg_priors are much different. Variables created through add_node are available for use in any node function, however variables passed through arg_priors are specific to only that node and the corresponding node_function.
-
Node functions must return a numpyro distribution object.
-
No cyclic behavior in the graph. But, you may use results from one node functions in another node functions. Allowing for infinite depth of nodes.
2. .create_model():
When the function .create_model() is called the class creates a graph representation of the nodes. It is able to do this through the node function’s arguments since the arguments are the parents of the respective nodes. If no node function is provided, the node has no parents. It then traverses the graph so that numpyro sample statements are defined in the correct order. Once finished the returned model can be used like any other numpyro model!
3. how to use plates
It can use plates using the plate argument in .add_node(). It takes in a tuple of plate objects allowing for infinitely nested plates.
from generative_model_variables import generative_model_variables
"""
Here I define the model variables object.
Small note:
If you need a constant value in a node function that you would like to pass in as a parameter.
You can pass in a dictionary to the model_args argument that maps from the name of a parameter to the value you would like to make available.
"""
variables = generative_model_variables(model_args={})
#plate
x_axis = numpyro.plate("data", len(data))
"""
Defining covariates
"""
#body size
variables.add_node(name='BS',
prior_distribution=dist.Categorical(logits=np.array([-2.2897760975840273, 0.9274377647576987, -1.4404764504491006])),
plate=(x_axis,))
#age at death
variables.add_node(name='age_at_death',
prior_distribution=dist.Categorical(logits=np.array([-4.239208520926547, -5.28994908412601, 3.846738437837947, -7.0859014343304585, -7.0859014343304585])),
plate=(x_axis,))
#trauma that breaks the skin
variables.add_node(name='trauma_br_skin',
prior_distribution=dist.Categorical(logits=np.array([-4.510859481371141, 0.09036250054334112, -0.2047254443224715, -4.0245010622075785])),
plate=(x_axis,))
#postmortem interval (i.e. how long the body as been dead)
variables.add_node(name='PMI',
prior_distribution=dist.Normal(2.1126173, 1.7476393),
plate=(x_axis,))
#clothing
variables.add_node(name='clothing',
prior_distribution=dist.Categorical(logits=np.array([0.39641516047,-1.623551043104,-1.166665718666])),
plate=(x_axis,))
"""
Here I loop over the decomposition characteristics since they
are all defined using the same information.
"""
#Decomposition characteristics
DECOMPS = ['livor_mortis_fixed','skin_slippage','marbling']
#node function
#parameters for this function must be defined either through add node or arg_priors
def function(clothing,beta_clothing,age_at_death,beta_age_at_death,trauma_br_skin,beta_trauma_br_skin,
PMI,BS,beta_BS,intercept,logpi_intercept):
logpi = PMI*(beta_clothing+beta_age_at_death+beta_trauma_br_skin\
+beta_BS+intercept)*PMI+logpi_intercept
return dist.Bernoulli(logits=logpi)
# These must match the names in the logpi function parameters
arg_priors = {
'beta_clothing':dist.Normal(np.array([0, 0, 0]), 0.5).to_event(1),
'beta_age_at_death': dist.Normal(np.array([0, 0, 0, 0, 0]), 0.5).to_event(1),
'beta_trauma_br_skin':dist.Normal(np.array([0, 0, 0, 0]), 0.5).to_event(1),
'beta_isHanging': dist.Normal(0, 0.5),
'beta_BS': dist.Normal(np.array([0, 0, 0]), 0.5).to_event(1),
'intercept': dist.Normal(-8,2),
'logpi_intercept': dist.Normal(-8,2)}
for node in DECOMPS:
variables.add_node(name=node,
node_function=function,
plate=(x_axis,),
arg_priors=arg_priors)
#create model
model = variables.create_model()
"""
Now use this model in the same way as any other numpyro model.
"""
Lastly, I have only included the differences in how I can define numpyro model functions in this post. However, I also implemented .fit() and .predict() steps into the class structure so that we could use sklearn with our model. That aspect of the code has been super helpful in getting cross-validation scores specifically. I’m happy to follow up with more information about that too!
I appreciate any and all criticism/suggestions for this code. This post is brief, so if there are any questions I am more than happy to try and answer them! I hope I was able to provide a decent explanation of how this was useful for my team and I!