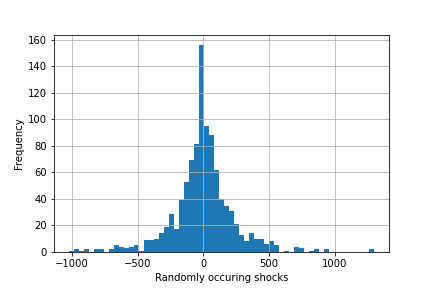
Sorry for the delay, but I am back working on this now. Could someone please give me some advice on the next steps?
Following the advice of @ fehiepsi above, I think I have implemented the model function correctly now and conditioned it on a set of observations (I modified it slightly so that initial guesses for two of the parameters can be provided):
import torch
import pyro
import pyro.distributions as dist
def random_shocks(scale_guess, b_guess):
scale = pyro.sample("scale", dist.Gamma(2, scale_guess)) # always positive
b = pyro.sample("b", dist.Gamma(2, b_guess)) # always positive
epsilon = pyro.param("epsilon", torch.tensor(0.01), constraint=constraints.interval(0, 1))
alpha = pyro.sample("alpha", dist.Bernoulli(epsilon))
alpha = alpha.long()
wp_dist = dist.Normal(torch.tensor([0.0, 0.0])[alpha], torch.tensor([scale, scale*b])[alpha])
measurement = pyro.sample('obs', wp_dist)
return measurement
def random_shocks_conditioned(scale_guess, b_guess, observations):
scale = pyro.sample("scale", dist.Gamma(2, scale_guess)) # always positive
b = pyro.sample("b", dist.Gamma(2, b_guess)) # always positive
epsilon = pyro.param("epsilon", torch.tensor(0.01), constraint=constraints.interval(0, 1))
alpha = pyro.sample("alpha", dist.Bernoulli(epsilon))
alpha = alpha.long()
wp_dist = dist.Normal(torch.tensor([0.0, 0.0])[alpha], torch.tensor([scale, scale*b])[alpha])
with pyro.plate("N", observations.shape[0]):
measurement = pyro.sample('obs', wp_dist, obs=observations)
My goal is to estimate the ‘scale’, ‘b’, and ‘epsilon’ parameters, and if possible, the sequence of binary random numbers alpha, given a sequence of observations. Is this possible with variational inference?
If I understand correctly, the next step is to create a parameterized guide function. In this case, I am thinking a Gaussian mixture distribution (with two components) would do the trick.
Which inference algorithm is appropriate here and is there an appropriate example script I could follow?