Hi the forum,
I have some specific questions for my model which I would like some feedback with. My NUTS and SVI results are wildly different. I’d like to know why, and how to remedy this.
The model, an extension of this example, is as follows:
reparam_config = {
k: LocScaleReparam(0)
for k in [
"alpha_s1",
"alpha_s2",
"alpha_s3",
"alpha_age_drift",
"beta_s1",
"beta_s2",
"beta_s3",
"beta_age_drift",
"xi",
"gamma_drift",
]
}
@numpyro.handlers.reparam(config=reparam_config)
def model(age, space, time, lookup12, lookup23, population, deaths=None):
N_s1 = len(np.unique(lookup12))
N_s2 = len(np.unique(lookup23))
N_s3 = len(np.unique(space))
N_age = len(np.unique(age))
N_t = len(np.unique(time))
N = len(population)
# plates
age_plate = numpyro.plate("age_groups", N_age, dim=-3)
space_plate = numpyro.plate("space", N_s3, dim=-2)
year_plate = numpyro.plate("year", N_t - 1, dim=-1)
# hyperparameters
sigma_alpha_s1 = numpyro.sample("sigma_alpha_s1", dist.HalfNormal(1.0))
sigma_alpha_s2 = numpyro.sample("sigma_alpha_s2", dist.HalfNormal(1.0))
sigma_alpha_s3 = numpyro.sample("sigma_alpha_s3", dist.HalfNormal(1.0))
sigma_alpha_age = numpyro.sample("sigma_alpha_age", dist.HalfNormal(1.0))
sigma_beta_s1 = numpyro.sample("sigma_beta_s1", dist.HalfNormal(1.0))
sigma_beta_s2 = numpyro.sample("sigma_beta_s2", dist.HalfNormal(1.0))
sigma_beta_s3 = numpyro.sample("sigma_beta_s3", dist.HalfNormal(1.0))
sigma_beta_age = numpyro.sample("sigma_beta_age", dist.HalfNormal(1.0))
sigma_xi = numpyro.sample("sigma_xi", dist.HalfNormal(1.0))
sigma_gamma = numpyro.sample("sigma_gamma", dist.HalfNormal(1.0))
# spatial hierarchy
with numpyro.plate("s1", N_s1, dim=-2):
alpha_s1 = numpyro.sample("alpha_s1", dist.Normal(0, sigma_alpha_s1))
beta_s1 = numpyro.sample("beta_s1", dist.Normal(0, sigma_beta_s1))
with numpyro.plate("s2", N_s2, dim=-2):
alpha_s2 = numpyro.sample(
"alpha_s2", dist.Normal(alpha_s1[lookup12], sigma_alpha_s2)
)
beta_s2 = numpyro.sample(
"beta_s2", dist.Normal(beta_s1[lookup12], sigma_beta_s2)
)
with space_plate:
alpha_s3 = numpyro.sample(
"alpha_s3", dist.Normal(alpha_s2[lookup23], sigma_alpha_s3)
)
beta_s3 = numpyro.sample(
"beta_s3", dist.Normal(beta_s2[lookup23], sigma_beta_s3)
)
beta_s3_cum = jnp.outer(beta_s3, jnp.arange(N_t))[jnp.newaxis, :, :]
# age
with age_plate:
alpha_age_drift_scale = jnp.pad(
jnp.broadcast_to(sigma_alpha_age, N_age - 1),
(1, 0),
constant_values=10.0, # pad so first term is alpha0, prior N(0, 10)
)[:, jnp.newaxis, jnp.newaxis]
alpha_age_drift = numpyro.sample(
"alpha_age_drift", dist.Normal(0, alpha_age_drift_scale)
)
alpha_age = jnp.cumsum(alpha_age_drift, -3)
beta_age_drift_scale = jnp.pad(
jnp.broadcast_to(sigma_beta_age, N_age - 1), (1, 0), constant_values=10.0
)[:, jnp.newaxis, jnp.newaxis]
beta_age_drift = numpyro.sample(
"beta_age_drift", dist.Normal(0, beta_age_drift_scale)
)
beta_age = jnp.cumsum(beta_age_drift, -3)
# age-space interaction
with age_plate, space_plate:
xi = numpyro.sample("xi", dist.Normal(0, sigma_xi))
# age-time random walk
with age_plate, year_plate:
gamma_drift = numpyro.sample("gamma_drift", dist.Normal(beta_age, sigma_gamma))
gamma = jnp.pad(jnp.cumsum(gamma_drift, -1), [(0, 0), (0, 0), (1, 0)])
# likelihood
latent_rate = alpha_s3 + alpha_age + beta_s3_cum + xi + gamma
with numpyro.plate("N", N):
mu_logit = latent_rate[age, space, time]
numpyro.sample("deaths", dist.Binomial(population, logits=mu_logit), obs=deaths)
rng_key = random.PRNGKey(args.rng_seed)
guide = autoguide.AutoLowRankMultivariateNormal(model)
optim = numpyro.optim.Adam(step_size=args.learning_rate)
svi = SVI(model, guide, optim, loss=Trace_ELBO())
svi_result = svi.run(rng_key, args.num_svi_steps, age, space, time, lookup12, lookup23, population, deaths)
Unfortunately due to the nature of the (identifiable health) data, I cannot show a reproducible example.
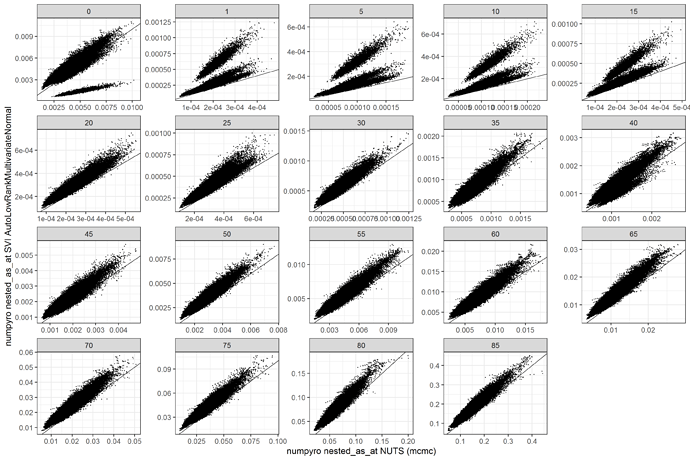
I am trying to get a similar posterior to my NUTS model, which converges well but takes about a day on GPU to run. My experiments with SVI have taken much less time (5-60 mins), but the posterior looks very different. Here is a comparison of the median estimates comparing numpyro to SVI with AutoLowRankMultivariateNormal. Clearly something is off.
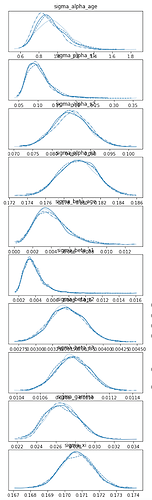
Also, the posterior estimates of the hyperparameters are off. First, the NUTS densities:
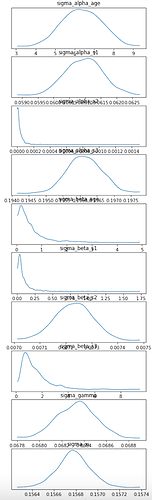
and these are the SVI densities
It is strange for
sigma_alpha_age to be so high when there is a HalfNormal(1.) prior.
I tried several different guides:
- AutoDiagonalNormal. Converged to a stable loss (100,000 svi steps, tried several learning rates settling on 0.1) presented the large
sigma_alpha_agevalue. - AutoLowRankMultivariateNormal. Converged to a stable loss (100,000 svi steps, tried several learning rates settling on 0.01) presented the large
sigma_alpha_agevalue and is well away from the NUTS posterior (see above). - AutoIAFNormal exhausted the 40GB GPU and did not run.
- AutoBNAFNormal led to ArrayMemoryError
- AutoDAIS had init_loss of 21386858981528199033645372704976119424776154659071… (+many more digits) and then led to nan loss. I tried different learning rates.
Let me know if there are any different strategies I should try, or if it is unlikely SVI will work for this model.
Cheers,
Theo