Hello Pyro experts,
I’m trying to implement a piecewise linear regression model. I’ve tried various variations on the model, but I haven’t been able to get good results. My approach is to sample change points from a Categorical and use a MaskedMixture with a linear regression as one component distribution, and another MaskedMixture as the other component (i.e. the MaskedMixture’s are nested as many times as there are pieces).
def piecewise_regression(x, y, n_pieces = 3):
"""
piecewise regression where pieces are connected and the last piece has slope 0
"""
N = x.shape[0]
lines = []
change_points = []
line_distributions = []
for piece in pyro.plate('pieces', n_pieces):
if piece == 0:
lines.append(sample_linear_regression())
line_distributions.append(dist.Normal(lines[-1][0]*x+lines[-1][1],lines[-1][2]))
change_point = sample_change_point(x, piece)
change_points.append(change_point)
elif piece < n_pieces-1:
mask = torch.arange(N) > change_points[-1]
slope, noise_std = sample_slope_and_noise_std(piece)
prev_slope = lines[-1][0]
prev_intercept = lines[-1][1]
intercept = (prev_slope*x[change_point]+prev_intercept)-slope*x[change_point]
lines.append((slope,intercept,noise_std))
prev_dist = line_distributions[-1]
line_distributions.append(dist.MaskedMixture(mask, prev_dist, dist.Normal(lines[-1][0]*x+lines[-1][1],lines[-1][2])))
# if all of x is masked, sample_change_point returns the last x
change_point = sample_change_point(x, piece)
change_point = min(change_point + change_points[-1], N)
change_points.append(change_point)
else:
noise_std = pyro.sample('noise_std_{}'.format(piece), dist.LogNormal(0.,1.))
intercept = lines[-1][0]*x[change_points[-1]]+lines[-1][1]
lines.append((intercept,noise_std))
prev_dist = line_distributions[-1]
last_mask = torch.arange(N) >= change_points[-1]
line_distributions.append(dist.MaskedMixture(last_mask, prev_dist, dist.Normal(lines[-1][0],lines[-1][1])))
with pyro.plate('N', N):
y = pyro.sample("obs", line_distributions[-1], obs=y)
return y, lines, change_points, line_distributions
def sample_change_point(x, piece, mask = None):
n_iter = x.shape[0]
if mask is None:
change_point_probs_prior = torch.ones(n_iter)
elif mask.any():
change_point_probs_prior = mask.float()
else:
return n_iter-1
change_point = pyro.sample('change_point_{}'.format(piece), dist.Categorical(probs=change_point_probs_prior),infer={'enumerate': 'parallel'})
return change_point
def sample_linear_regression():
slope = pyro.sample('slope', dist.Normal(0.,1.))
intercept = pyro.sample('intercept', dist.Normal(0.,1.))
noise_std = pyro.sample('noise_std', dist.LogNormal(0.,1.))
return slope, intercept, noise_std
def sample_slope_and_noise_std(piece):
slope = pyro.sample('slope_{}'.format(piece), dist.Normal(0.,1.))
noise_std = pyro.sample('noise_std_{}'.format(piece), dist.LogNormal(0.,1.))
return slope, noise_std
The guide has all the same distributions as the model: https://pastebin.com/sYmCpR6B
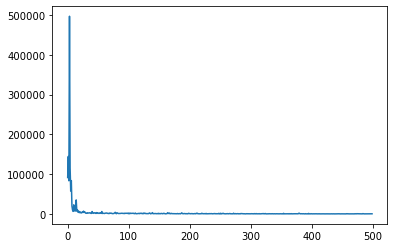
I want to enumerate the change points out with TraceEnum_ELBO, though it looks like that won’t scale very well with more change points. But even with just one change point, I don’t get a good fit, even though the elbo converges:
pyro.clear_param_store()
optim = pyro.optim.Adam({"lr": 0.05})
elbo = TraceEnum_ELBO(max_plate_nesting=1, num_particles=10)
svi = SVI(piecewise_regression, piecewise_regressionGuide, optim, loss=elbo)
losses = []
for i in range(500):
losses.append(svi.step(x,y,2))
plt.plot(losses)
pred = pyro.infer.predictive.Predictive(pyro.poutine.uncondition(piecewise_regression),guide=piecewise_regressionGuide,num_samples=100)
fit = pred(x,y,2)
fit_obs_mean = fit['obs'].mean(0).detach().numpy()
fit_obs_std = fit['obs'].std(0).detach().numpy()
%matplotlib inline
plt.plot(x,y)
#plt.plot(x,slope_fit*x.numpy()+intercept_fit)
#plt.plot(x,fit_obs)
plt.errorbar(x,fit_obs_mean,yerr=fit_obs_std)
plt.axis('equal');
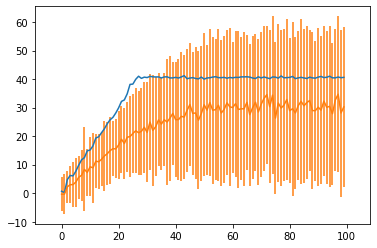
What can I do to make the fit more accurate? Is there a better way to implement this model? I’ve tried having Dirichlet priors on the change points, and multiple restarts (which helps a little, but best of 10 restarts still isn’t great).
Thanks
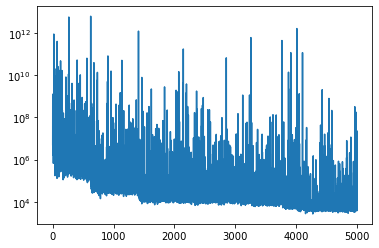 (that’s the AutoDiagonalNormal loc gradient)
(that’s the AutoDiagonalNormal loc gradient)