Hi, There is an example of Gaussian Mixture model in Pyro using TraceEnum_ELBO(Gaussian Mixture Model — Pyro Tutorials 1.8.4 documentation). I understand the idea is to use enumeration to marginalise the discrete variable called “assignment”.
Apart from this, I also notice that TraceGraph_ELBO can also deal with discrete variable and it seems that it doesn’t need the enumeration strategy. Therefore I want to try TraceGraph_ELBO instead. However, It seems that I failed to get the correct result. I’m assuming that one possible reason can be that I didn’t write the guide() correctly.
Below are my code and result. Could you please give me some advice? Thank you!
import torch
import os
import numpy as np
from torch.distributions import constraints
import pyro
import pyro.distributions as dist
from pyro.optim import Adam
from pyro.infer import SVI, TraceGraph_ELBO
smoke_test = ('CI' in os.environ)
assert pyro.__version__.startswith('1.8.1')
from pyro.distributions.torch import Multinomial, Beta, Dirichlet, Beta, Categorical, MultivariateNormal, Uniform
K = 2 # Fixed number of components.
data = torch.tensor([0., 1., 10., 11., 12.])
def model(data):
# Global variables.
weights = pyro.sample('weights', dist.Dirichlet(0.5 * torch.ones(K)))
scale = pyro.sample('scale', dist.LogNormal(0., 2.))
with pyro.plate('components', K):
locs = pyro.sample('locs', dist.Normal(0., 10.))
with pyro.plate('data', len(data)):
# Local variables.
assignment = pyro.sample('assignment', dist.Categorical(weights))
pyro.sample('obs', dist.Normal(locs[assignment], scale), obs=data)
def guide(data):
weights = pyro.sample('weights', dist.Dirichlet(0.5 * torch.ones(K)))
# scale
scale_para_loc = pyro.param('scale_para_loc', torch.tensor(1.),constraint=constraints.positive)
scale = pyro.sample('scale', dist.LogNormal(scale_para_loc, 2.))
# locs
locs_para_loc = pyro.param('locs_para_loc', torch.tensor(5.))
locs_para_scale = pyro.param('locs_para_scale', torch.tensor(1.),constraint=constraints.positive)
with pyro.plate('components', K):
locs = pyro.sample('locs', dist.Normal(locs_para_loc, locs_para_scale))
with pyro.plate('data', len(data)):
# Local variables.
assignment = pyro.sample('assignment', dist.Categorical(weights))
pyro.clear_param_store()
optim = pyro.optim.Adam({'lr': 0.1, 'betas': [0.8, 0.99]})
elbo = TraceGraph_ELBO()
svi = SVI(model, guide, optim, loss=elbo)
losses = []
for i in range(501 if not smoke_test else 2):
loss = svi.step(data)
losses.append(loss)
if i % 10 == 0:
print("ELBO at iter i = "+str(i),loss)
from pyro.infer import Predictive, SVI, Trace_ELBO, TraceGraph_ELBO
num_samples = 10
predictive = Predictive(model, guide=guide, num_samples=num_samples)
graph_samples = predictive(data)
graph_samples['assignment']
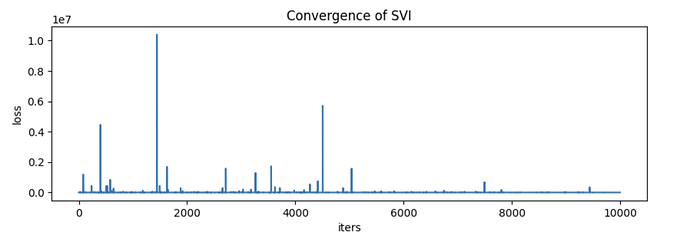
However, the ELBO doesn’t convergence at all. And the estimation results of the variable “assignment” doesn’t make any sense. Below are the 10 samples of “assignment” :
tensor(
[[0, 0, 0, 0, 0],
[0, 0, 1, 0, 0],
[1, 1, 1, 1, 1],
[1, 1, 1, 1, 1],
[1, 1, 1, 1, 1],
[0, 1, 1, 0, 1],
[0, 0, 1, 0, 1],
[1, 0, 0, 0, 1],
[1, 1, 1, 1, 1],
[1, 1, 1, 1, 1]])
The samples of “locs” also looks wrong:
tensor(
[[4.0562, 5.4942],
[4.2813, 6.4910],
[9.0555, 2.7641],
[6.8727, 6.9697],
[8.4839, 0.8544],
[2.1208, 8.5360],
[6.4476, 7.4402],
[5.7618, 3.1934],
[2.0187, 4.2319],
[2.9078, 4.8222]])
Thank you in advance for your help!