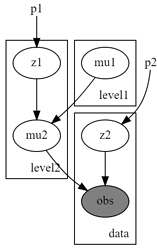
I’m having trouble with a hierarchical mixture model, whose posterior inference is performed with autoguide and naive enumeration (I’m not confident about whether I’m enumerating properly). I’m showing a toy example that should be reproduceable, and raises the same error.
def toy_model(data=None, n1=3, n2=5):
p1 = pyro.param("p1", torch.randn(n1).exp(), constraint=constraints.simplex)
p2 = pyro.param("p2", torch.randn(n2).exp(), constraint=constraints.simplex)
with pyro.plate("level1", n1):
mu1 = pyro.sample("mu1", Normal(0, 1))
with pyro.plate("level2", n2):
z1 = pyro.sample("z1", Categorical(p1), infer={'enumerate': 'parallel'})
mu2 = pyro.sample("mu2", Normal(mu1[z1], 0.1))
with pyro.plate("data", N):
z2 = pyro.sample("z2", Categorical(p2), infer={'enumerate': 'parallel'}) #, infer={'enumerate': 'parallel'}
pyro.sample("obs", Normal(mu2[z2], 0.01), obs=data)
model = toy_model
And the graph illustrates the hierarchical structure:
optim = ClippedAdam({'lr': 0.1, 'betas': [0.9, 0.999], 'lrd': 0.999})
elbo = pyro.infer.TraceEnum_ELBO(num_particles=4, max_plate_nesting=1)
def train(num_iterations, losses=[]):
for j in tqdm(range(num_iterations)):
loss = svi.step(data) / N
losses.append(loss)
return losses
def initialize(seed):
global global_guide, svi
pyro.set_rng_seed(seed)
pyro.clear_param_store()
global_guide = pyro.infer.autoguide.AutoDelta(
poutine.block(model, hide=['z1', 'z2']),
)
svi = SVI(model, global_guide, optim, loss=elbo)
return svi.loss(model, global_guide, data) / N
loss, seed = min((initialize(seed), seed) for seed in tqdm(range(100)))
initialize(seed)
print('seed = {}, initial_loss = {}'.format(seed, loss))
losses = []
losses = train(200, losses=losses)
ELBO loss converges:
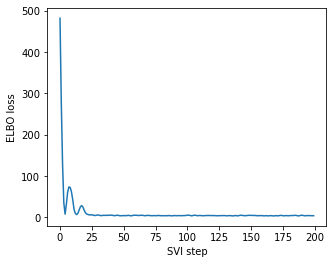
Here comes the problem: when I’m directly following the patterns here to perform MAP inference for discrete sites
guide_trace = poutine.trace(global_guide).get_trace(data) # record the globals
trained_model = poutine.replay(model, trace=guide_trace) # replay the globals
def classifier(data, temperature=0): #random assignment if temperature=1
inferred_model = infer_discrete(trained_model, temperature=temperature,
first_available_dim=-2) # avoid conflict with data plate
trace = poutine.trace(inferred_model).get_trace(data)
return trace.nodes["z2"]["value"]
classifier(data)
I got an error
ValueError: Error while packing tensors at site 'mu2':
Invalid tensor shape.
Allowed dims: -1
Actual shape: (3, 5)
Try adding shape assertions for your model's sample values and distribution parameters.
Trace Shapes:
Param Sites:
p1 3
p2 5
Sample Sites:
mu1 dist 3 |
value 3 |
log_prob 3 |
z1 dist 5 |
value 3 1 |
log_prob 3 5 |
mu2 dist 3 5 |
value 5 |
log_prob 3 5 |
Trace Shapes:
Param Sites:
Sample Sites:
The infer_discrete and classifier worked well for programs with only one discrete site, and after checking the shape, I thought the error should be related with enumeration. How should I deal with this issue?