Main code here
def init_svi(self, X: DeviceArray, *, lr: float, **kwargs):
"""Initialize the SVI state
Args:
X: input data
lr: learning rate
kwargs: other keyword arguments for optimizer
"""
self.optim = self.optim_builder(lr, **kwargs)
self.svi = SVI(self.model, self.guide, self.optim, self.loss)
svi_state = self.svi.init(self.rng_key, X)
if self.svi_state is None:
self.svi_state = svi_state
return self
def _fit(self, X: DeviceArray, n_epochs) -> float:
@jit
def train_epochs(svi_state, n_epochs):
def train_one_epoch(_, val):
loss, svi_state = val
svi_state, loss = self.svi.update(svi_state, X)
return loss, svi_state
return lax.fori_loop(0, n_epochs, train_one_epoch, (0., svi_state))
loss, self.svi_state = train_epochs(self.svi_state, n_epochs)
return float(loss / X.shape[0])
code mainly from GitHub - FlorianWilhelm/bhm-at-scale: 🪜 Bayesian Hierarchical Models at Scale .
I got nan while trainning
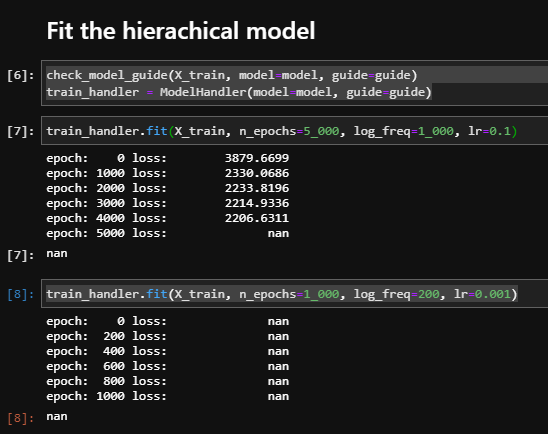
I tried to debug it, but all value I got is like this
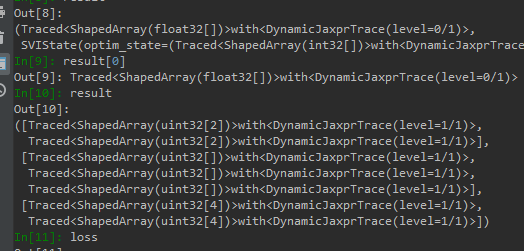
how do I debug it ?
I think you can remove jit and convert lax.fori_loop to a for loop. Alternatively, you can use some utilities control_flow_prims_disabled and fori_loop to avoid compiling your function.
Do you have any good resouces to quickly understand the key points of jit and lax.fori_loop using in numpyro ?
You can find a nice exposition here. In NumPyro, we use jit + for loop if we want to display a progress bar or print out some useful information during the loop. Otherwise, using fori_loop is much faster for small body_fn. Does this make sense to you?
If you want to debug, just use the usual for loop and remove jit. The control_flow_prims_disabled utility converts fori_loop to a for loop, which helps for debugging if you don’t want to rewrite the _fit method. If you want to keep jit in _fit method, then you can use jax.disable_jit function to disable jit. In summary, you can use numpyro.util.fori_loop instead of jax.lax.fori_loop in _fit method, then use the context managers numpyro.util.control_flow_prims_disabled together with jax.disable_jit in the notebook to debug. You can also use numpyro.enable_validation() to trigger some warnings if parameters get wrong values.
Your are so kind , thank you very much !
Thanks for all these hints with respect to debugging. Maybe a special section in NumPyro’s docs listing all these hints would be a great addition.
Besides what you listed, I would like to add:
import sys
import warnings
from IPython.core import ultratb
warnings.simplefilter("error") # turn warnings to errors
# activate debugger on error
sys.excepthook = ultratb.FormattedTB(mode="Verbose", color_scheme="Linux", call_pdb=1)
This helped me a lot when values outside of the support of a distribution got evaluated. By default this only triggers a UserWarning:
UserWarning: Out-of-support values provided to log prob method. The value argument should be within the support.
warnings.warn('Out-of-support values provided to log prob method. '
With the code above those are turned to errors and in order to find the culprit, a debugger is launched to find the name of the distribution and location in your source code.
Also thanks for pointing out numpyro.util.fori_loop @fehiepsi. Would you recommend to always use numpyro’s fori_loop in case you want some iteration of some kind or is JAX’s scan preferred? Reading this JAX scan vs fori_loop discussion it feels like scan is the better fori_loop but in our case we don’t really neet the differentiability of scan but how is it with respect to performance?
3 Likes
how is it with respect to performance?
I think the performance should be similar (I am not sure). If you don’t need differentiation and collect something, it is probably simpler to write code in fori_loop pattern, and probably easier to debug. Thanks for the tips by the way, triggering an error would be very helpful for debugging!

