Thanks for this tip, it helped me identify a surprising defect in my implementation. When I first tested with a Normal distribution, things looked plausible:
def model():
numpyro.sample('x', dist.Normal(0,1))
kernel = Metropolis(model, step_size=.1)
mcmc = MCMC(kernel, num_warmup=0, num_samples=500_000, thinning=500)
mcmc.run(rng_key, init_params={'x':0.0})
posterior_samples = mcmc.get_samples()
plt.hist(posterior_samples['x'], bins=50)
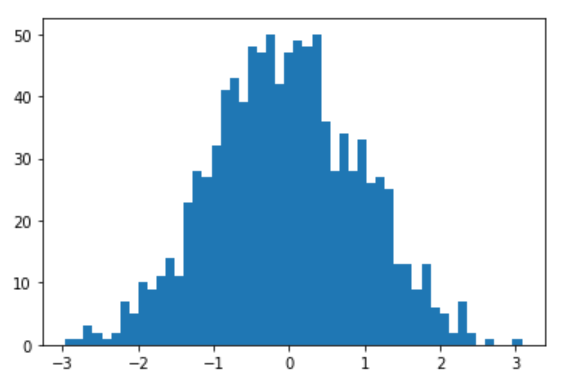
But when I tried with a simple constrained distribution instead (Uniform), things are clearly not right:
def model():
x = numpyro.sample('x', dist.Uniform(0,1))
kernel = Metropolis(model, step_size=.1)
mcmc = MCMC(kernel, num_warmup=0, num_samples=500_000, thinning=500)
mcmc.run(rng_key, init_params={'x':0.0})
posterior_samples = mcmc.get_samples()
plt.hist(posterior_samples['x'], bins=50)
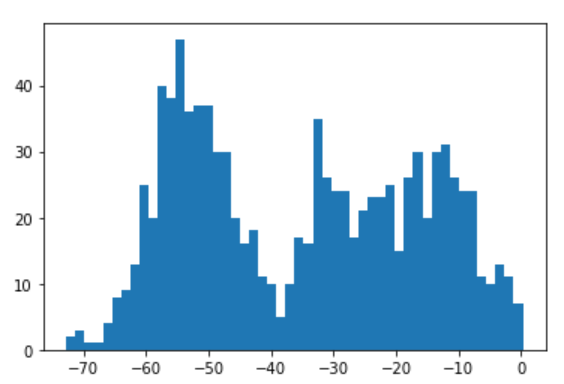
(Note that these samples are not constrained to the interval [0,1].)
I followed the example code you pointed me to and got something that stays within the Uniform’s support. It ended up needing a lot more code in the init method than my first attempt, though. Am I over-complicating things? Again any feedback is welcome!
class Metropolis(MCMCKernel):
def __init__(self, model, step_size=0.1):
self._model = model
self._step_size = step_size
@property
def sample_field(self):
return "z"
@property
def default_fields(self):
return ("z",)
def init(self, rng_key, num_warmup, init_params, model_args, model_kwargs):
assert rng_key.ndim == 1, "only non-vectorized, for now"
init_params_2, potential_fn, postprocess_fn, model_trace = util.initialize_model(
rng_key,
self._model,
# init_strategy=self._init_strategy,
dynamic_args=True,
model_args=model_args,
model_kwargs=model_kwargs,
)
self._potential_fn = potential_fn
self._postprocess_fn = postprocess_fn
_, unravel_fn = ravel_pytree(init_params)
def _make_log_prob_fn(potential_fn, unravel_fn):
def log_prob_fn(x):
return -potential_fn(unravel_fn(x))
return log_prob_fn
self._log_prob_fn = _make_log_prob_fn(
potential_fn(*model_args, **model_kwargs), unravel_fn
)
return MetState(init_params_2.z, rng_key)
def postprocess_fn(self, args, kwargs):
if self._postprocess_fn is None:
return identity
return self._postprocess_fn(*args, **kwargs)
def sample(self, state, model_args, model_kwargs):
rng_key, key_proposal, key_accept = random.split(state.rng_key, 3)
z_flat, unravel_fn = ravel_pytree(state.z)
z_proposal = dist.Normal(z_flat, self._step_size).sample(key_proposal)
z_proposal_dict = unravel_fn(z_proposal)
log_pr_0 = self._log_prob_fn(z_flat)
log_pr_1 = self._log_prob_fn(z_proposal)
accept_prob = jnp.exp(log_pr_1 - log_pr_0)
z_new = jnp.where(dist.Uniform().sample(key_accept) < accept_prob, z_proposal, z_flat)
return MetState(unravel_fn(z_new), rng_key)
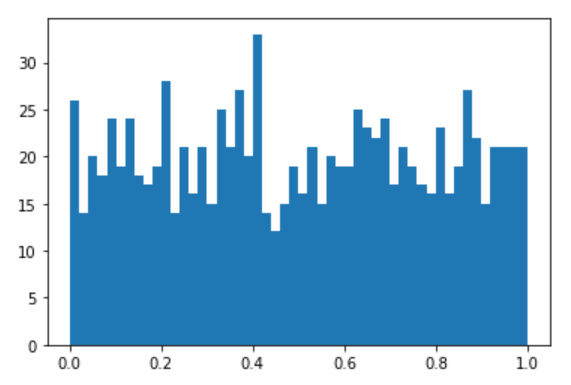
I’ve put the code for the class and a self-contained example run in this gist so that it has line numbers and revision control.


