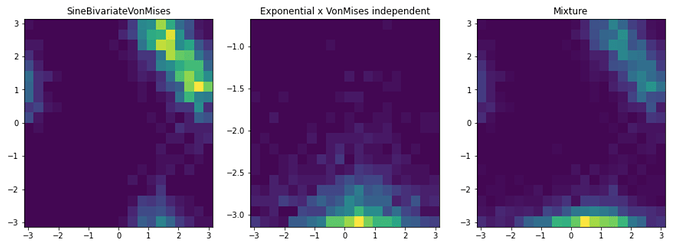
Hey, I would like to implement a GeneralMixture model on two-dimensional data.
My models work just fine when using the builtin multidimensional distributions like MultivariateNormal or BivariateVonMises. But I also would like to use just two univariate distributions that are independent of each other, in addition to that (so have both in one model).
Q: What I don’t know though is how I would model a bivariate distribution (dist_IND) composed out of two independent random variables, that sample from univariate builtin distributions (like a Normal (dist_Ix and an Exponential (dist_Iy)).
I guess what would work is to just manually implement a bivariate distribution that samples from the two univariate distributions individually and puts them together. But maybe there is sth builtin, that I did not find so far?
What I found was the Independent class, but from what I understood this is only used to basically reduce the correlation matrix of a MultivariateNormal distribution.
Thanks in advance.
P.S. This is my first post, so sorry if this is not the best description, but don’t hesitate to ask for any further information, I am more than willing to do so.
Some not working code:
# shape of data would be: (num_samples, 2)
def general_mixture_independent(data, num_components):
weights = numpyro.sample('weights', dist.Dirichlet(jnp.ones(num_components)))
# multivariate builtin distribution
mu_MVN = numpyro.sample('mu_MVN', dist.MultivariateNormal(jnp.zeros(2), jnp.eye(2)))
dist_MVN = dist.MultivariateNormal(mu_MVN, jnp.eye(2))
# two univariate builting distributions
# x-dimension
mu_x = numpyro.sample('mu_x', dist.Normal(0, 1))
sigma_x = numpyro.sample('sigma_x', dist.HalfNormal(1))
dist_Ix = dist.Normal(mu_x, sigma_x)
# y-dimension
lambda_y = numpyro.sample('lambda_y', dist.HalfNormal(3))
dist_Iy = dist.Exponential(lambda_y)
# put together -> that's basically what I don't know how to do
# and it of course does not work like that
dist_IND = jnp.column_stack([dist_Ix, distIy])
mixture = dist.MixtureGeneral(dist.Categorical(probs=weights), [dist_MVN, dist_IND], obs=data)