I noticed that if I run
x2_grid = jnp.linspace(-20,70,1000)
sigma= jnp.exp(dist.Uniform(low=0, high=50).log_prob(x2_grid))
I obtain this plot
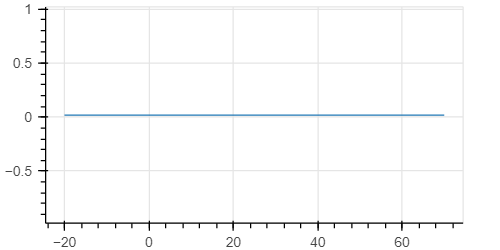
while if I run
x2_grid = jnp.linspace(-20,70,1000)
sigma= jnp.exp(dist.Uniform(low=0, high=50, validate_args=True).log_prob(x2_grid))
I obtain what I expected, namely:
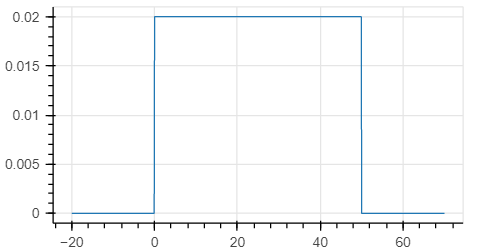
In the documentation it says that this parameter " Whether to enable validation of distribution parameters and arguments to .log_prob method." But it is not clear to me what it means, and why it returns a constant value if I run it without the validation.
with validate_args=True the parameters of the distribution and the sample values passed to log_prob are checked to see if they satisfy required constraints. for example the scale of a Normal distribution needs to be positive and a sample value passed to LogNormal.log_prob must be positive.
in the case of the uniform distribution the log_prob is log(1/(high - low)) in support and -inf out of support. so in the validate_args=True case extra checks are done so that you correctly get -inf out of support. in the validate_args=False case these checks are skipped (for additional speed) under the assumption that you’re on the hook for passing in sample values that are actually in support and results are not guaranteed to be correct (or even not nans) if you pass in sample values that are out of support
1 Like

