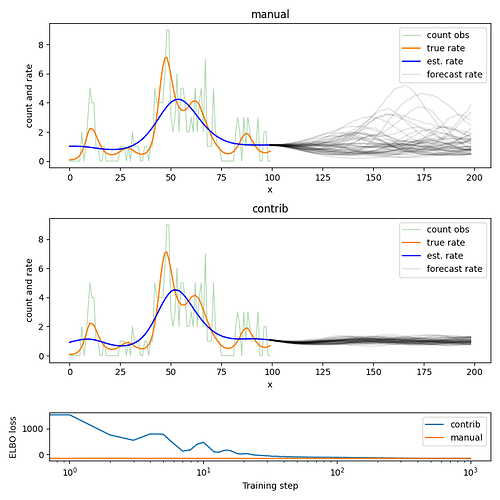
I’m learning Pyro, and trying to get my head around the global hidden state, in particular within the Gaussian Process pyro.contrib. I’ve reduced my model into a simple latent Poisson rate model to show below.
The following code shows how I’m trying to approach the problem with the pyro.contrib.gp which I can’t get working, and I’ve also written a GP manually (switched with use_contrib argument) that does work.
The variance and lengthscale don’t seem to be fitting via pyro.contrib.gp. I’m fitting with an AutoDelta for MAP estimation, but the same results occur when using other guides such as AutoMutlivariateNormal.
def model(x: torch.Tensor,
count_obs: Optional[torch.Tensor] = None,
x_pred: Optional[torch.Tensor] = None,
use_contrib=False):
n = len(x)
x = x.unsqueeze(-1)
jitter = torch.tensor(1e-4)
# Priors for RBF kernel lengthscale and variance.
lam = pyro.sample('lam', dist.Gamma(torch.tensor(14.), torch.tensor(1.)))
tau = pyro.sample('tau', dist.Gamma(torch.tensor(2.), torch.tensor(4.)))
if use_contrib:
# Covariance matrix. (contrib: this does not work)
kern = gp.kernels.RBF(1)
kern.variance = tau
kern.lengthscale = lam
cov_x = kern.forward(x)
else:
# Covariance matrix. (manual: this works)
cov_x = tau * torch.exp(-(x - x.t())**2/lam**2)
# Sample latent rate.
cov_x = cov_x + jitter * torch.eye(n)
lograte = pyro.sample('lograte', dist.MultivariateNormal(torch.zeros(n), cov_x))
# Observations.
with pyro.plate('x', n):
pyro.sample('count_obs', dist.Poisson(rate=lograte.exp()), obs=count_obs)
# Predictions.
if x_pred is not None:
if use_contrib:
# Get a GP for the estimated latent rate. (contrib: this does not work)
kern = gp.kernels.RBF(1)
kern.variance = tau
kern.lengthscale = lam
gp_pred = gp.models.GPRegression(x.squeeze(), lograte, kern, jitter=jitter)
gp_pred.noise = torch.tensor(0.) # No noise since this is the latent rate.
lograte_pred_mean, lograte_pred_cov = gp_pred.forward(x_pred, full_cov=True)
else:
# Get a GP for the estimated latent rate. (manual: this works)
x_pred = x_pred.unsqueeze(-1)
cov_x_pred = tau * torch.exp(-(x - x_pred.t())**2/lam**2)
cov_x_pred_pred = tau * torch.exp(-(x_pred - x_pred.t())**2/lam**2)
cov_x_chol = torch.linalg.cholesky(cov_x)
alpha = torch.cholesky_solve(lograte.unsqueeze(-1), cov_x_chol)
v = torch.linalg.solve_triangular(cov_x_chol, cov_x_pred, upper=False)
lograte_pred_mean = (cov_x_pred.T @ alpha).squeeze()
lograte_pred_cov = cov_x_pred_pred - v.t() @ v
# Sample predicted latent rate.
lograte_pred_cov = lograte_pred_cov + jitter * torch.eye(len(x_pred))
lograte_pred = pyro.sample('lograte_pred', dist.MultivariateNormal(lograte_pred_mean, lograte_pred_cov))
# Sample predicted counts.
with pyro.plate('x_pred', len(x_pred)):
pyro.sample('count_pred', dist.Poisson(rate=lograte_pred.exp()))