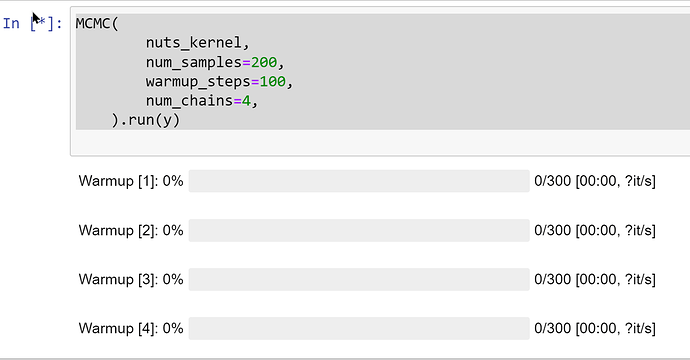
I was trying to follow up the example of MCMC with and LKJ prior and make some modification to let it runnable in the Jupyter Notebook, here is the code segment I am running
import argparse
import torch
import pyro
import pyro.distributions as dist
from pyro.infer.mcmc import NUTS
from pyro.infer.mcmc.api import MCMC
def model(y):
d = y.shape[1]
N = y.shape[0]
options = dict(dtype=y.dtype, device=y.device)
# Vector of variances for each of the d variables
theta = pyro.sample("theta", dist.HalfCauchy(torch.ones(d, **options)))
# Lower cholesky factor of a correlation matrix
concentration = torch.ones(
(), **options
) # Implies a uniform distribution over correlation matrices
L_omega = pyro.sample("L_omega", dist.LKJCholesky(d, concentration))
# Lower cholesky factor of the covariance matrix
L_Omega = torch.mm(torch.diag(theta.sqrt()), L_omega)
# For inference with SVI, one might prefer to use torch.bmm(theta.sqrt().diag_embed(), L_omega)
# Vector of expectations
mu = torch.zeros(d, **options)
with pyro.plate("observations", N):
obs = pyro.sample("obs", dist.MultivariateNormal(mu, scale_tril=L_Omega), obs=y)
return obs
y = torch.randn(500, 5).to(dtype=torch.double)
nuts_kernel = NUTS(model, jit_compile=True, step_size=1e-5)
MCMC(
nuts_kernel,
num_samples=200,
warmup_steps=100,
num_chains=4,
).run(y)
but it seems that the MCMC. run keeps running without any progress, which is shown in the following screenshot